FROM CANCER METHYLOME ATLAS TO BLOOD-BASED SCREENING: A TARGETED MULTI-OMICS APPROACH FOR CFDNA MULTI-CANCER EARLY DETECTION
2023-01-17
Utilizing blood cfDNA methylation for multi-cancer early detection (MCED) is ushering in a transformative era in cancer screening. Early attempts, such as Grail's Galleri test, have demonstrated in multiple clinical trials the capability to detect DNA signals associated with numerous deadly cancers before symptoms emerge, while also providing cancer signal origin (CSO) prediction. However, due to reliance on techniques like bisulfite conversion that cause significant DNA damage, Galleri's current sensitivity for cancer detection—particularly for early-stage cancers—remains relatively low. Additionally, differences in cancer spectra and incidence between China and the US, possible epigenetic etiological variations, and Galleri's relatively high cost reduce its potential for cost-effectiveness within China's healthcare system. Given its enormous potential, developing an MCED kit tailored to the Chinese cancer spectrum—with significantly improved sensitivity and substantially reduced cost—has become an important objective. Here, using our proprietary, bisulfite-free, high-sensitivity DNA methylation detection technology (Carbernet), we constructed a cancer DNA methylation atlas specific to the Chinese population and designed, built, and iteratively validated a targeted hybrid-capture methylation panel covering multi-omic markers for MCED. The database encompasses whole-genome DNA methylation and hydroxymethylation analyses of 1,263 paired tumor and adjacent normal tissues across nine high-incidence cancers in China, 1,795 plasma cfDNA samples from cancer patients, and 780 plasma samples from healthy individuals, spanning various clinical stages, ages, and sexes. The latest version of the MCED kit integrates multi-omic features—including methylation, fragmentomics, methylation haplotype patterns, and DNA mutations—into a unified predictive model, achieving significantly improved sensitivity early-stage cancer detection across a total of 6,296 plasma samples tested. This work establishes the largest cancer DNA methylome reference to date and demonstrates the clinical feasibility of a targeted, multi-omics approach for cfDNA-based MCED and tissue-of-origin prediction. Future efforts will focus on expanding the database to include additional cancer types and further iterating the assay to enhance both detection performance and cancer type coverage.
A Targeted Multi-Omics Approach for cfDNA Multi-Cancer Early Detection

Cabernet: A Proprietary, High-sensitivity DNA Methylation Profiling Technology

Enzymatic conversion enables high detection sensitivity of cfDNA methylation signals released by early-stage cancers
Tailored for plasma cell‑free DNA, preserving double‑stranded DNA integrity and retaining fragmentation information
Cancer DNA Methylome Atlas based on China’s Cancer Spectra
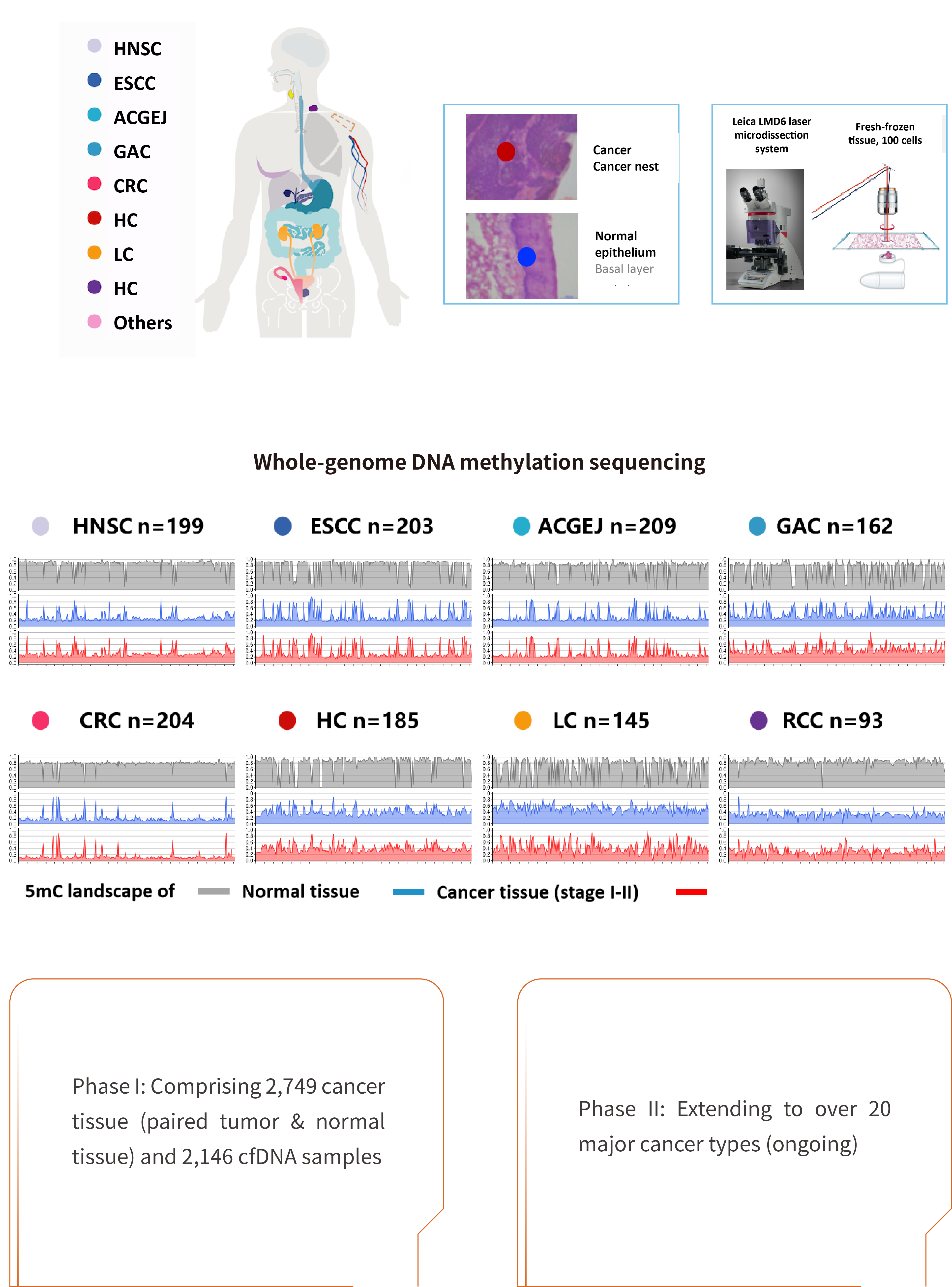
Iterative optimization of targeted probes for multi-cancer early detection

Automation and cost reduction for future large-scale MCED deployment
End‑to‑end “blood sample to data” automation on an assembly line, processing 20,000–50,000 samples per month
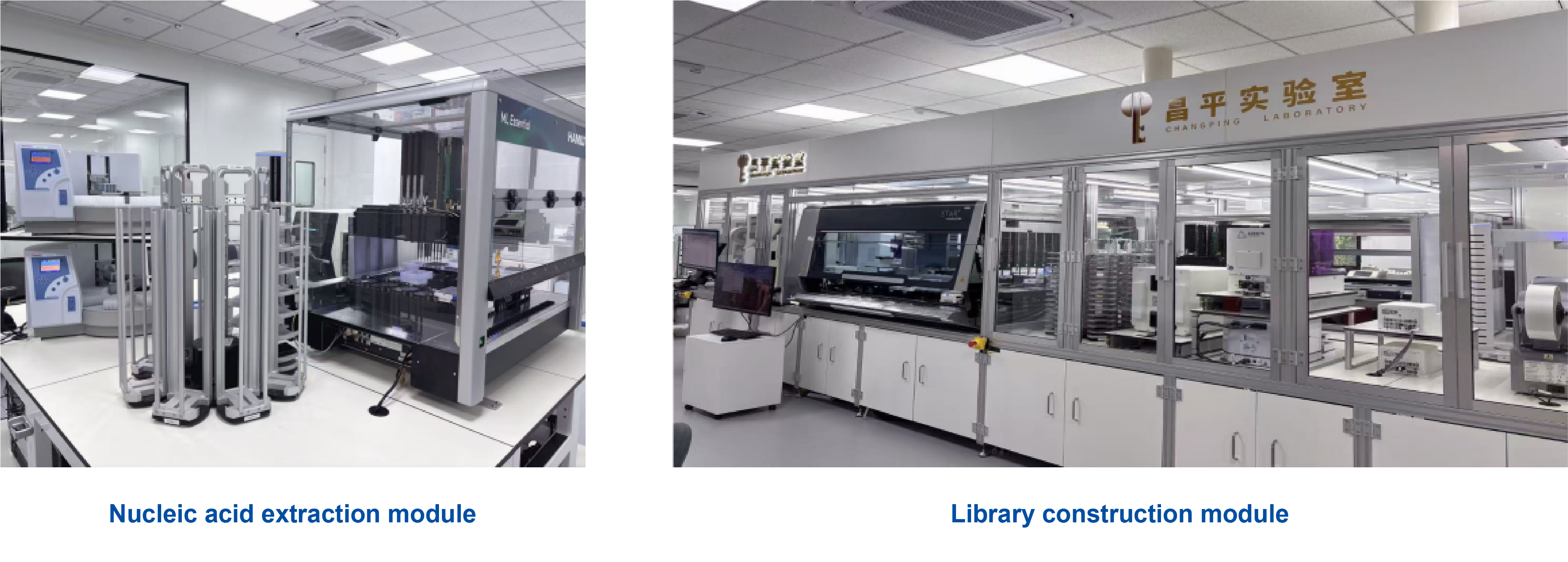

Domestication of reagents and sequencing platform, expected to be priced at 1/10 of competing MCED kit.